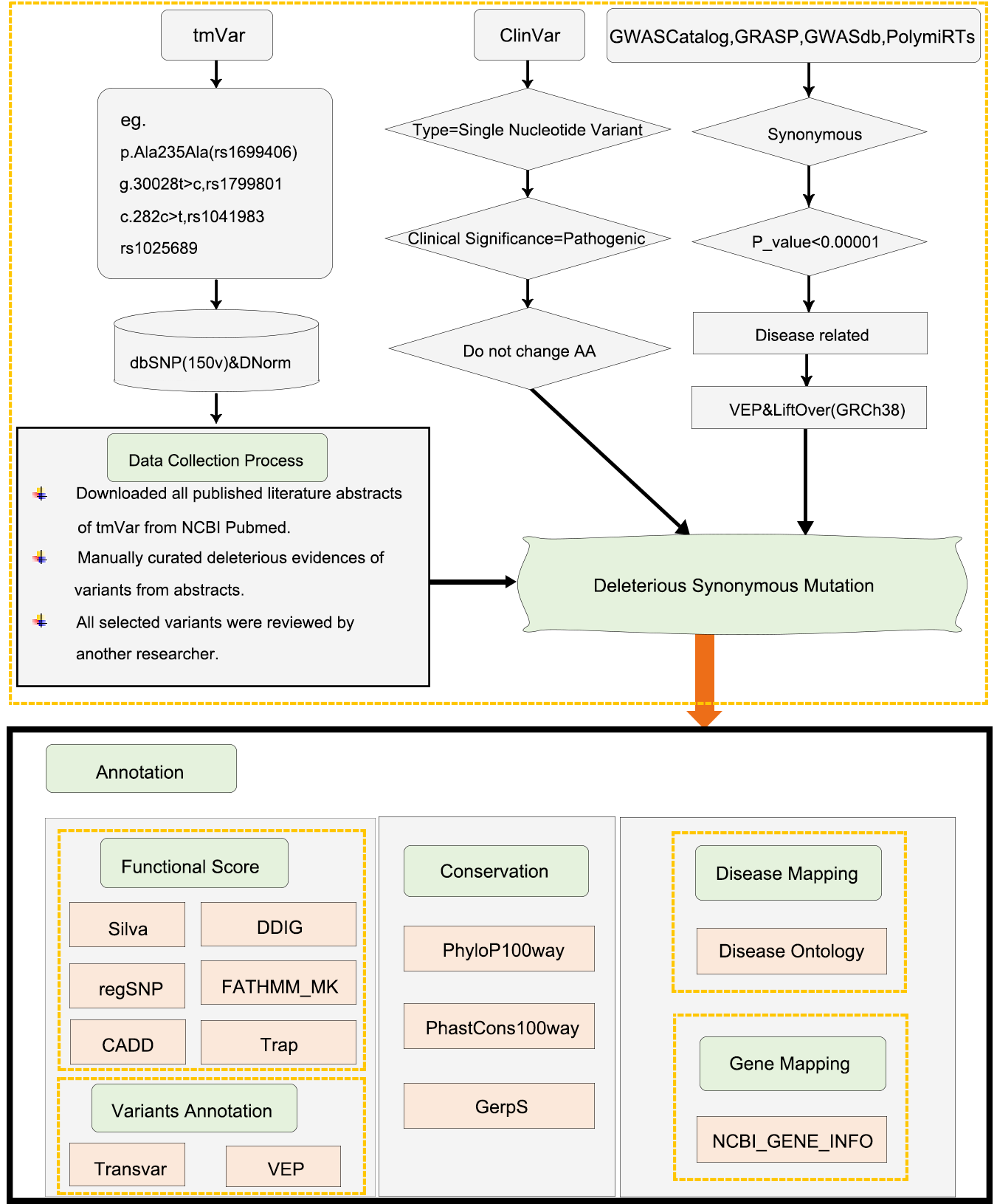
Overview
Synonymous mutation is a form of single nucleotide mutation which replaces one base to another in the coding sequence that does not alter amino acids in the translated protein due to the degeneracy genetic code. Because of synonymous mutation does not change the protein primary structure, it has been considered nonfunctional for a long time. So it sometimes called 'silent mutation'. But recent studies suggesting that such synonymous mutations play an important role in human phenotype by affecting transcript splicing, mRNA stability, RNA secondary structure, the biding site of transcription factor binding and microRNA, codon optimality and protein expression and so on.
dbDSM (Database of Deleterious Synonymous Mutation) is an integrated database that collect multiple sources relate to deleterious synonymous mutations (DSMs). Here, we present the updated dbDSMv2.0 database, which provides detail information about DSM with phenotypes, including position information, disease, gene and functional score etc. We also add the source of tmVar to dbDSMv2.0 which is a text mining approach for extracting a wide range of sequence variants in both protein and gene levels (e.g. substitution, deletion, etc) in biomedical literature. We intend to collect accurate information used for bio-research and make it accessible to all users freely.
Citation:
Pengbo Wen, Peng Xiao, and Junfeng Xia*, dbDSM: a manually curated database for deleterious synonymous mutations. Bioinformatics, 2016, 32(12): 1914-1916.