-
About dbCPM
-
Web Browser Requirements
-
Evidence System
-
Usage
- 1. Browse
- 2. Search
- 3. Download
- 4. Contact
-
Database Summary
-
Version
About dbCPM
For developing computational methods to predict the effects of cancer mutations and for evaluating the false positive rates of cancer driver mutation detection efforts, benchmark datasets containing both positive (driver mutations) and negative (passenger mutations) datasets are needed. While some recently developed datasets for cancer driver mutations are available for this purpose, benchmark sets for cancer passenger mutations have largely been missing. Therefore, we developed a comprehensive literature-based database of Cancer Passenger Mutations (dbCPM). In the current release, dbCPM contains 941 experimentally supported and 978 putative passenger mutations derived by manual curation of literature. All mutation descriptions include the position information (gDNA, cDNA, and protein levels), specific diseases, evidence sentences as well as evidence levels (in vivo, in vitro, or putative). The database can facilitate the development and evaluation of computational methods for predicting the effects of cancer mutations.
To obtain genomic positions, we used TransVar (http://bioinformatics.mdanderson.org/transvarweb/) through entering genes and their associated cDNA changes and mapped them to the results of the longest possible transcripts. Genomic positions that failed to match the canonical cDNA at the specified site were substituted by a dot (.). To acquire standard disease terminology, we mapped the related disease onto Disease Ontology (http://www.disease-ontology.org/).
Please cite the paper, if you are using the information in the database:
Yue Z, Zhao L, Xia J. dbCPM: a manually curated database for exploring the cancer passenger mutations. Briefings in bioinformatics, 2018. doi.org/10.1093/bib/bby105.
Web Browser Requirements
The dbCPM requires a modern web browser with JavaScript and cookies enabled. To view the complex details, pop-ups must not be blocked. The following browsers have been thoroughly tested with dbCPM:
- Mozilla Firefox, version 4 or above
- Internet Explorer, versions 9 or above
- Chrome, version 5 or above
Evidence System
Levels of evidence for mutations in dbCPM.
| Level No. | Details |
|---|---|
| Level 1 (in vivo) |
Alteration is regarded as a passenger based on evidence from functional experiments in vivo. |
| Level 2 (In vitro) |
Alteration is regarded as a passenger based on evidence from functional experiments In vitro. |
| Level 3 (Putative) |
Alteration is a putative passenger based on evidence such as a high recurrence frequency in healthy controls, an unimportant location in protein and so on. |
Usage
In dbCPM, we tried to make it powerful and convenient to be used. This Usage is prepared for the online service. The dbCPM provides the browse function, search function and download function at present.
Capitalised titles correspond to column headings in the web page tables:
- DISEASE: disease terminology in Disease Ontology
- DOID: Disease Ontology identifier
- GENE: The official gene symbol
- Transcript: Transcript stabel ID from Ensembl Genome Browser
- TYPE: missense, synonymous, nonsense, indel (insertion or deletion) or noncoding
- HIGHEST LEVEL: The highest evidence level of variants across specified both disease and gene
1. Browse
You can select one or more of the four options listed in the browse area (Disease, Gene, Mutation and Evidence). The Mutation option only can be available after a gene is selected.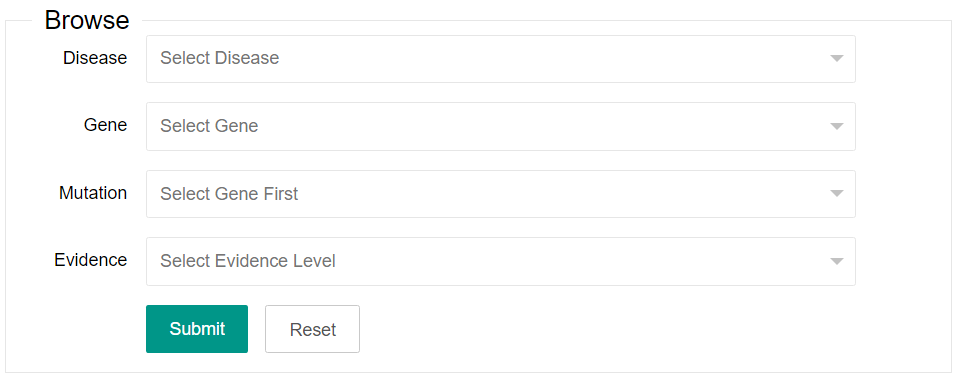
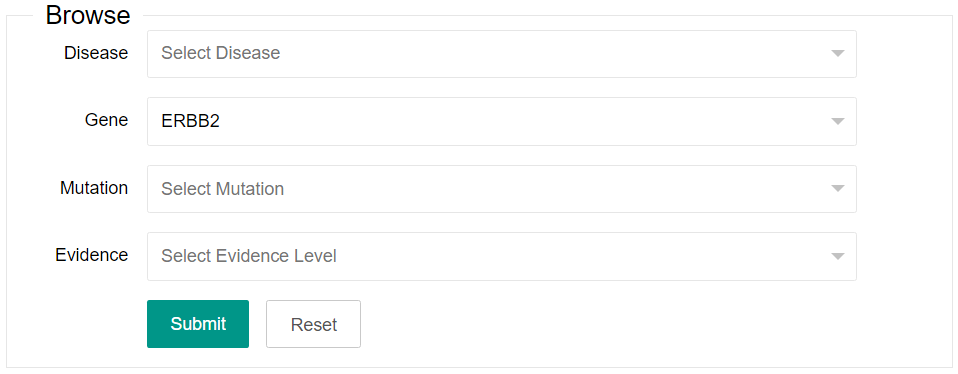
2. Search
You can input one keyword to search the dbCPM. The search fields include disease name, gene name or mutation (gDNA, cDNA or protein positions). Only the GRCh37 genome assembly is supported.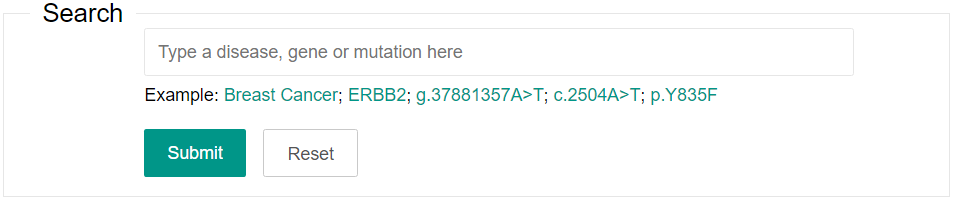
3. Download
We provide the option to download the full database. If you'd like to download it, please click to download the data for each level.
4. Contact
I have a few questions which are not listed above, how can I contact the authors of dbCPM?Please contact Dr. Junfeng Xia (Email: jfxia@ahu.edu.cn) for details.
Database Summary (current version)
(A) Database statistics
(B) Type distribution
| Gene | Mutation | Disease | PMID | Entry | Level 1 (in vivo) |
Level 2 (in vitro) |
Level 3 (putative) |
| 161 | 1919 | 43 | 312 | 3813 | 157 | 1112 | 2544 |
| Missense | Synonymous | Nonsense | Indel | Noncoding | Total |
| 1673 | 127 | 6 | 24 | 89 | 1919 |
Indel: insertion or deletion.
Version
The version 1.1 was released on June 8th, 2018.
The version 1.0 was released on January 4th, 2018.
The version 1.0 was released on January 4th, 2018.